Ambiscript permits the rapid derivation of complementary sequences
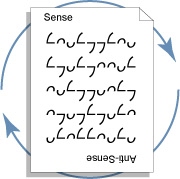
Since important genetic information can reside on either strand of the DNA double helix, it is often necessary to derive the sequence which complements that reported in genetic databases. In the traditional IUPAC nation, this is accomplished by first reversing the order of the characters in the sequence and then replacing each with the symbol for its complement. Ambiscript character symmetries eliminate the need for this two-step error-prone process.
It turns out that when complementary nucleotides are encoded by pairs of rotationally-related ambigrams, one can complement a sequence of any size by simply rotating the text 180°1. The term ambigram was first used by Douglas Hofstadter to describe symbols which convey the same or different meanings when viewed in different orientations2.
Since Ambiscript uses ambigrams to represent complementary nucleotides, it is possible to derive the complement of any Ambiscript-encoded sequence by rotating the text 180°, as shown in the figure. The rotational symmetries of this ambigraphic nucleic acid notation also make it easy to identify genetic palindromes as discussed in the following section.
1 Rozak DA. 2006. The practical and pedagogical advantaes of an ambigraphic nucleic acid notation. Nucleosides Nucleotides, and Nucleic Acids 25:807-813. (PMID: 16898419)
2 Hofstadter DR. 1985. Metamagical Themas: Questioning for the Essence of Mind and Patern. Basic Books, NY.